SPIKEPIPE: A metagenomic pipeline for the accurate quantification of eukaryotic species occurrences and abundances using DNA barcodes or mitogenomes
New publication by Yinqiu Ji, Tea Huotari, Tomas Roslin, Niels Martin-Schmidt, Jiaxin Wang, View ORCID Profile Douglas Yu, Otso Ovaskainen
Abstract:
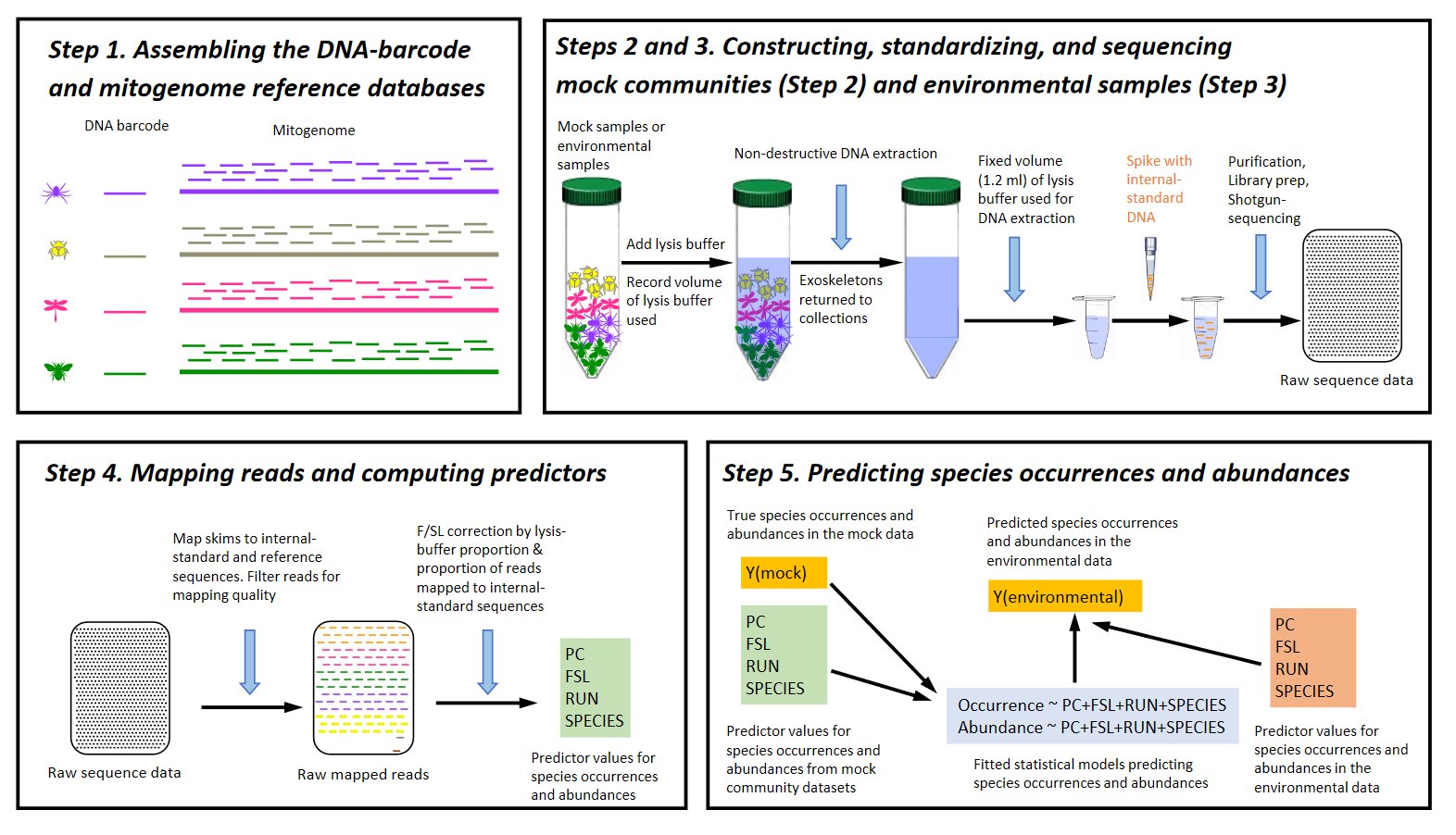
The accurate quantification of eukaryotic species abundances from bulk samples remains a key challenge for community description and environmental biomonitoring. We resolve this challenge by combining shotgun sequencing, mapping to reference DNA barcodes or to mitogenomes, and three correction factors: (1) a percent-coverage threshold to filter out false positives, (2) an internal-standard DNA spike-in to correct for stochasticity during sequencing, and (3) technical replicates to correct for stochasticity across sequencing runs. This pipeline achieves a strikingly high accuracy of intraspecific abundance estimates from samples of known composition (mapping to barcodes R2=0.93, mitogenomes R2=0.95) and a high repeatability across environmental-sample replicates (barcodes R2=0.94, mitogenomes R2=0.93). As proof of concept, we sequence arthropod samples from the High Arctic systematically collected over 17 years, detecting changes in species richness, abundance, and phenology using either barcodes or mitogenomes. SPIKEPIPE provides cost-efficient and reliable quantification of eukaryotic communities, with direct application to environmental biomonitoring.
bioRxiv. doi.org/10.1101/533737
